Pharmacophore Generation & Screening
1. Foreword
Pharmacophore refers to a combination of a series of spatial and electronic characteristics that can match the characteristics of a specific biological target , and the molecules that match this characteristic can excite or block the biological response of the target . Pharmacophore-based virtual screening is often used to screen potential new drugs.
The Hermite platform's Pharmacophore Generation & Screening module provides pharmacophore generation and virtual screening capabilities. With the massive virtual compound library built into the Hermite platform, you can quickly screen potential new drugs.
2. Usage
2.1 Entrance
The general menu bar on the left Function → Virtual Screening → Pharmacophore Generation & Screening.
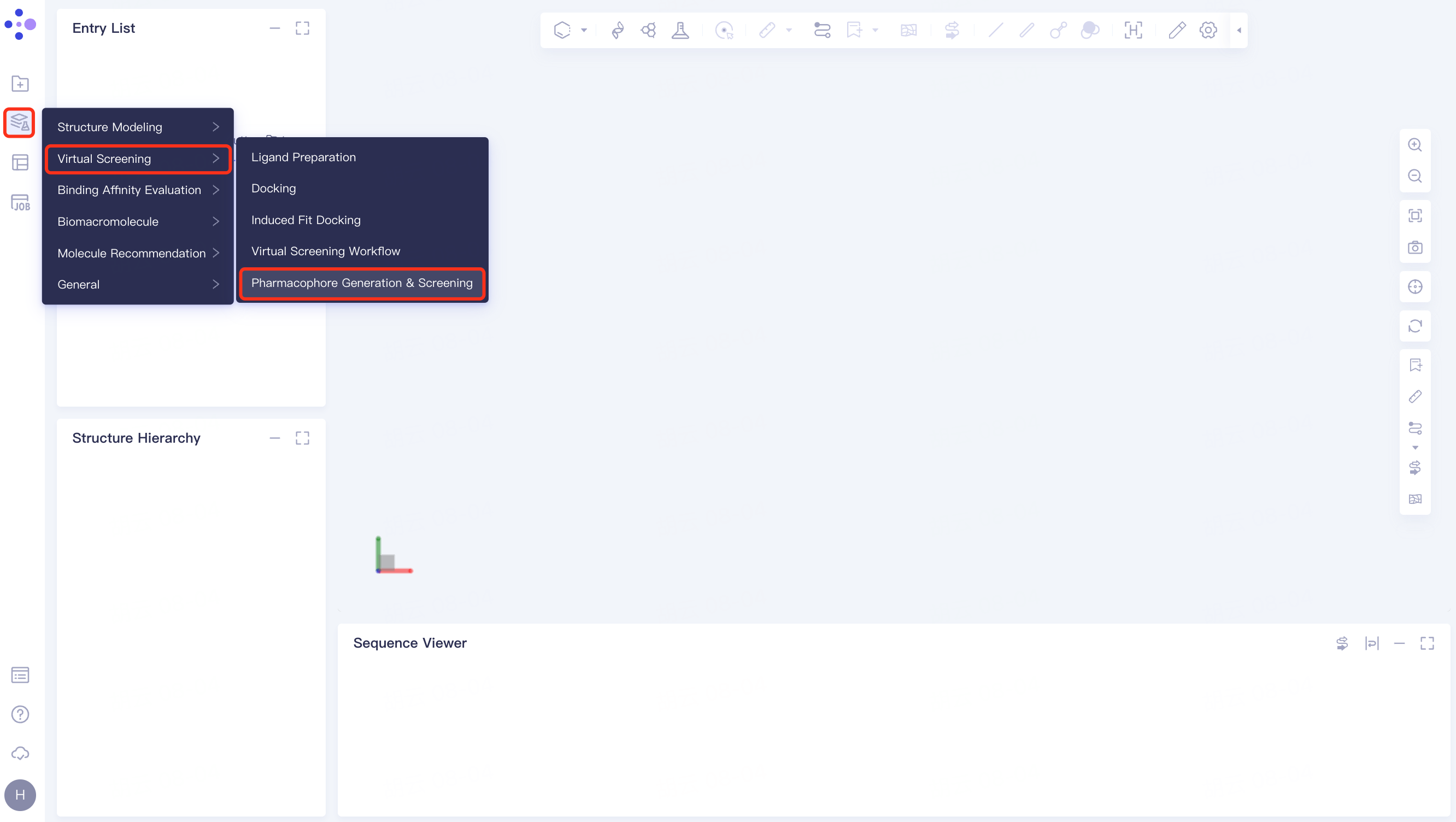
The following interface pops up:
Click "Yes" to enter the steps of 2.2, and click "No" to enter the steps of 2.3.
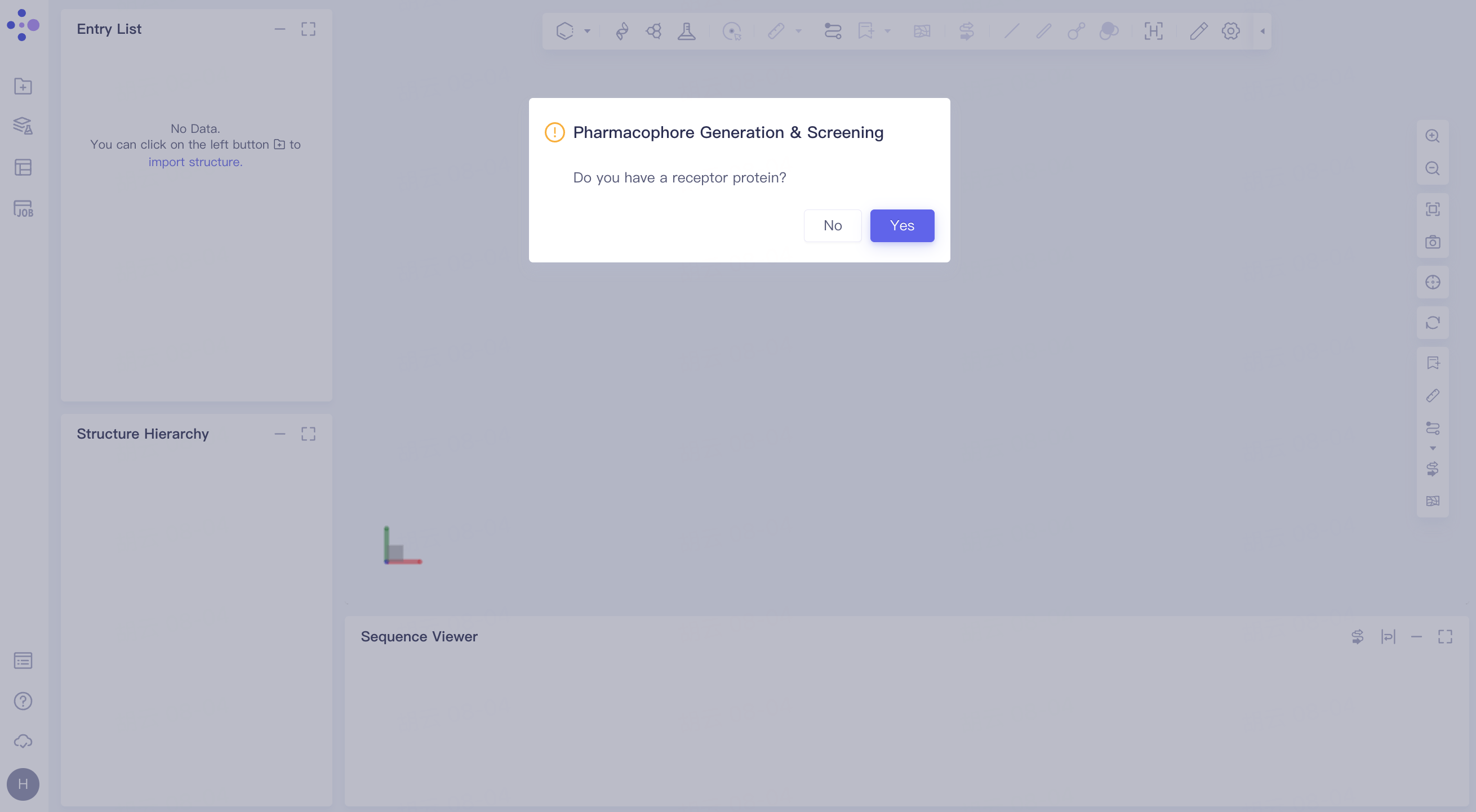
2.2 Pharmacophore generation and screening with receptor protein
2.2.1 selector protein
There are three ways:
(1)From 3D:
Click the Select from 3D Workspace checkbox → pop up the Select Structure interface → Select the required protein structure in the Structure Hierarchy box on the left side of the interface/Select the protein structure in the 3D Workspace window → Select Structure interface to display the selected protein name, click OK.
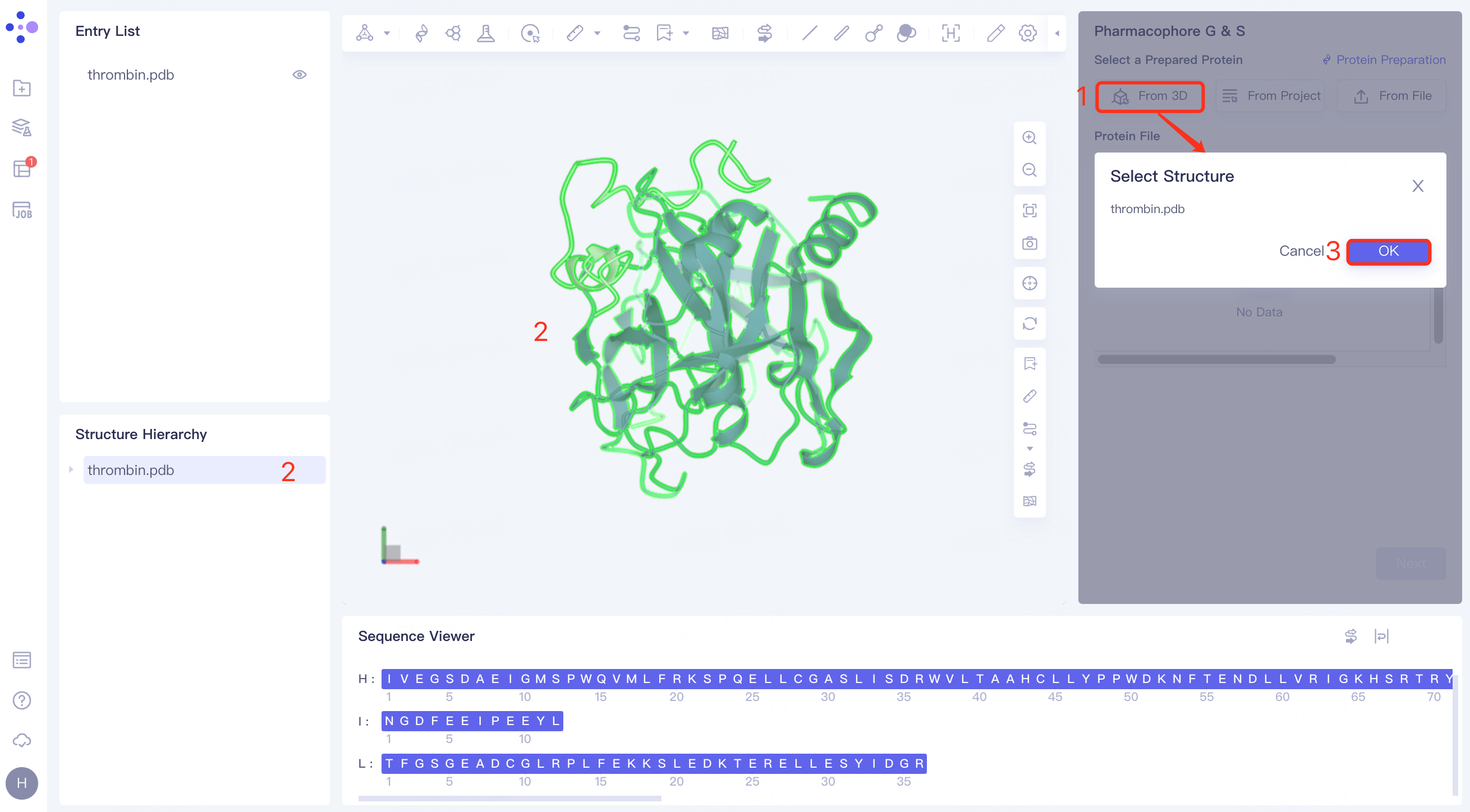
(2)From Project:
Click the Select from Project checkbox → pop up the Select from Project interface → select the desired protein structure → click OK.
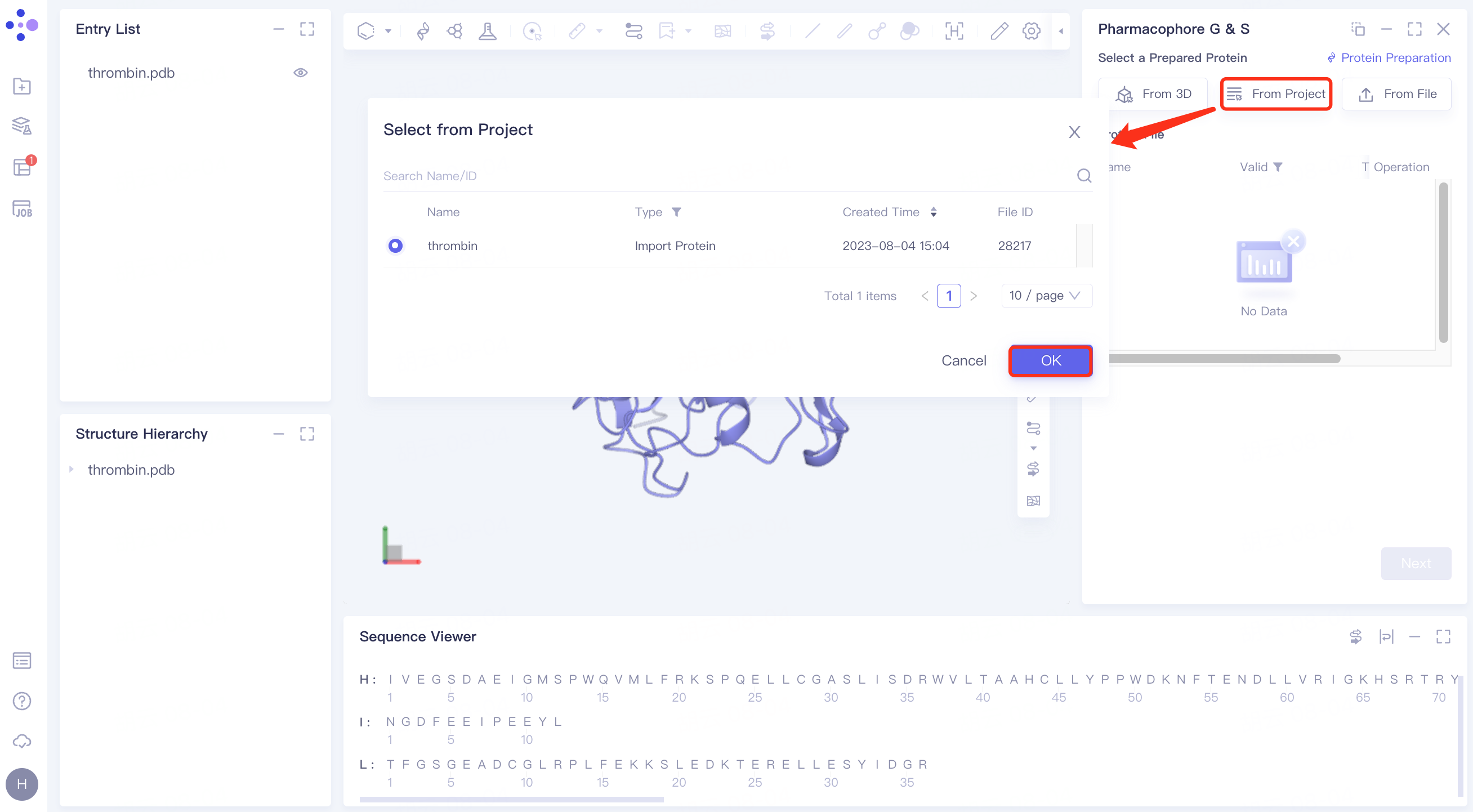
(3)From File:
Click the Select File checkbox → Select the desired protein file (.pdb format file is supported) from the local folder and upload it.
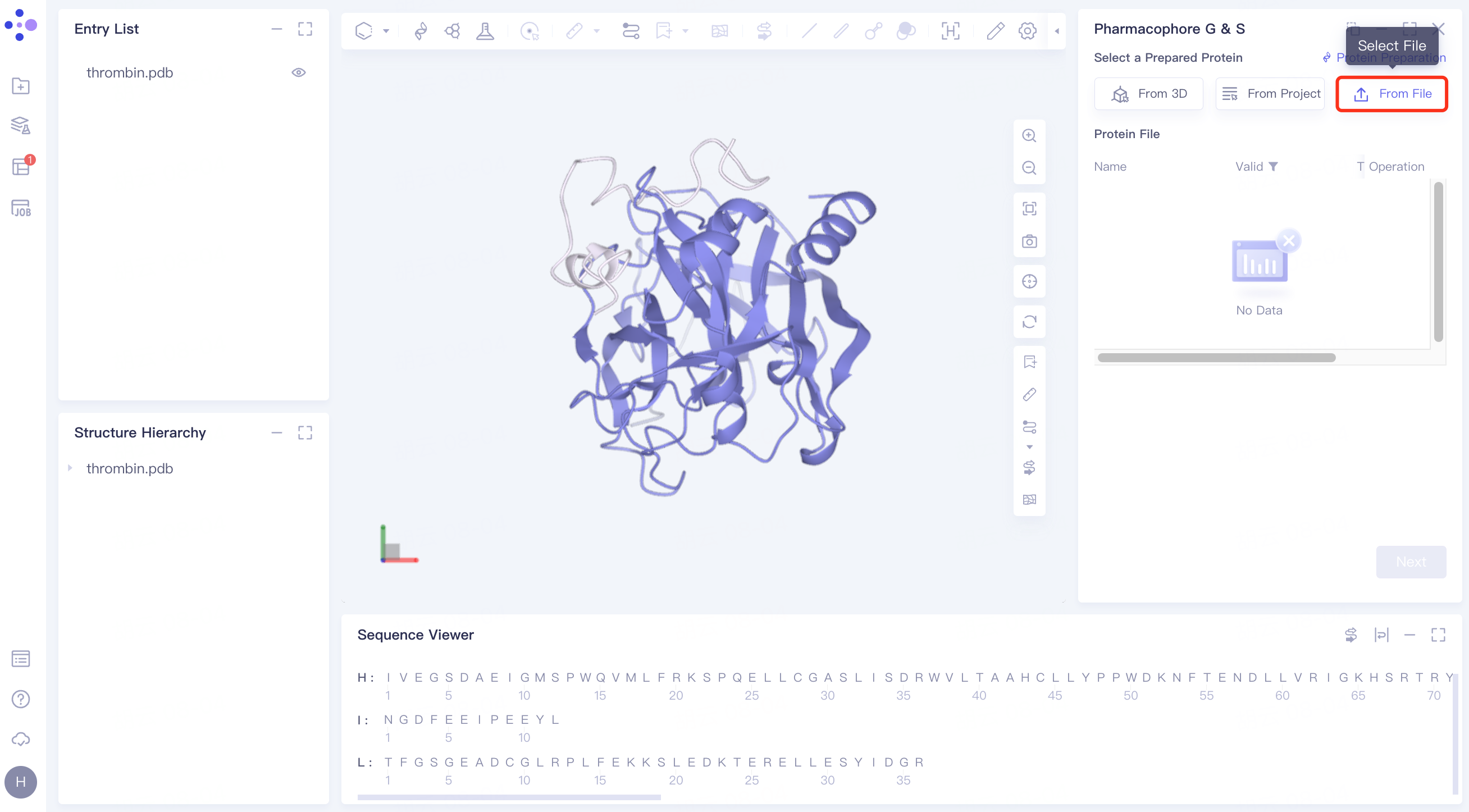
2.2.2 upload reference ligand
There are three ways:
(1)Select from 3D
Click Select from 3D → Pop-up Select Ligands interface → Select the desired ligand structure in the Structure Hierarchy box on the left side of the interface/Select the ligand structure in the 3D Workspace window → Select the Ligands interface to display the selection
For the ligand name, click OK.
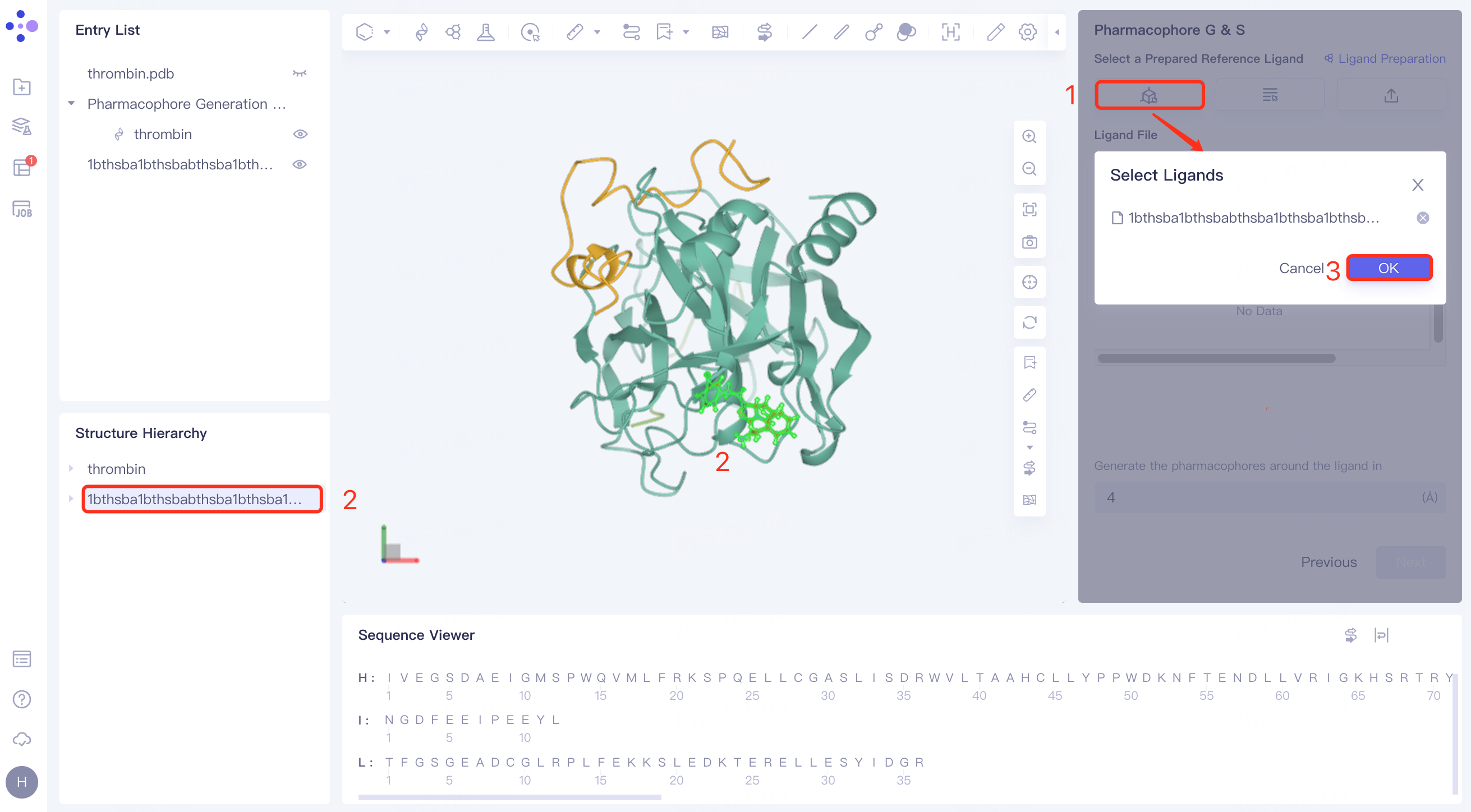
(2)Select from Project
Click the Select from Project checkbox → pop-up Select from Project interface → select the desired ligand structure → click OK.
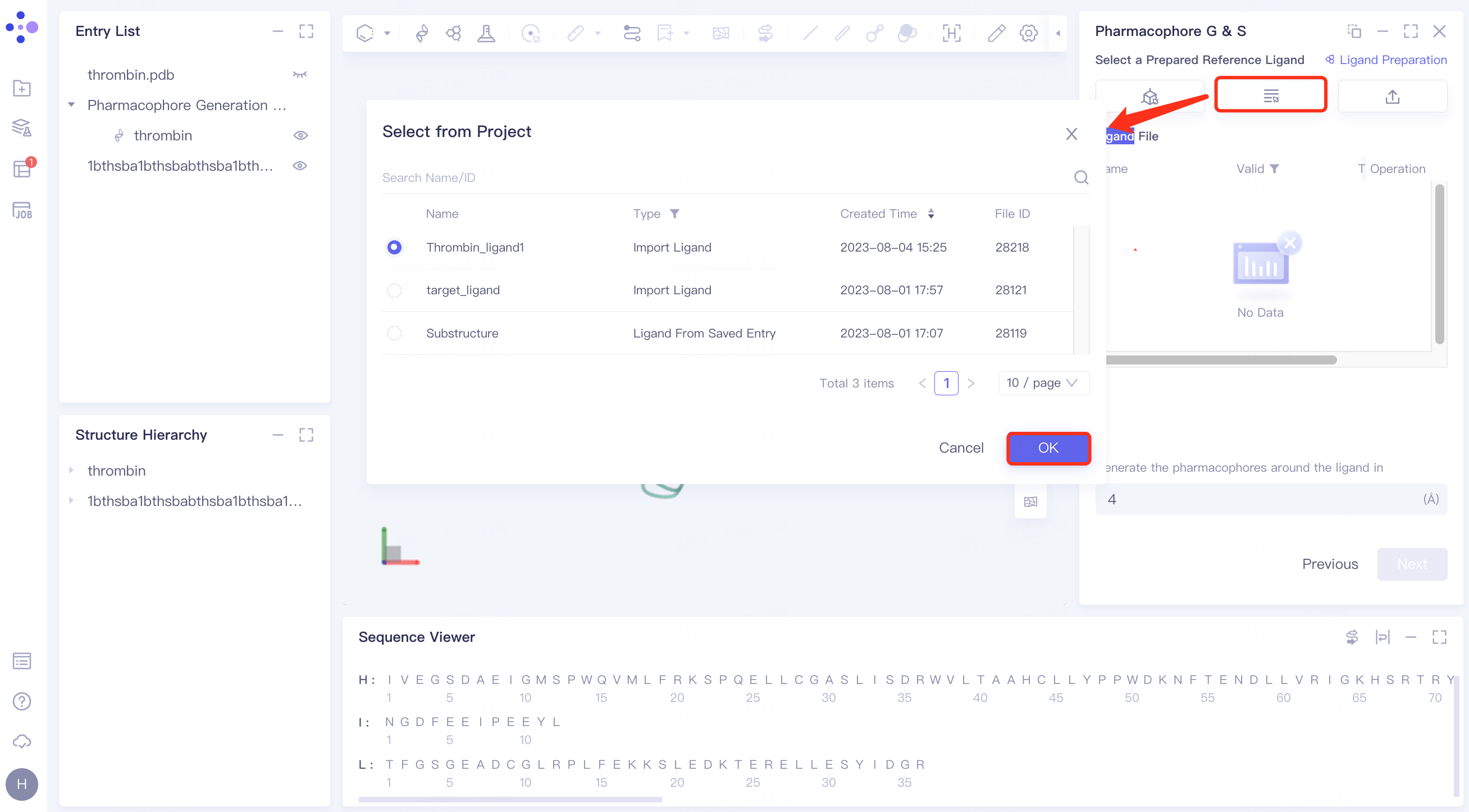
(3)Select File
Click the Select File checkbox → Select the desired ligand file (.mol and .sdf file formats are supported) from the local folder and upload it.
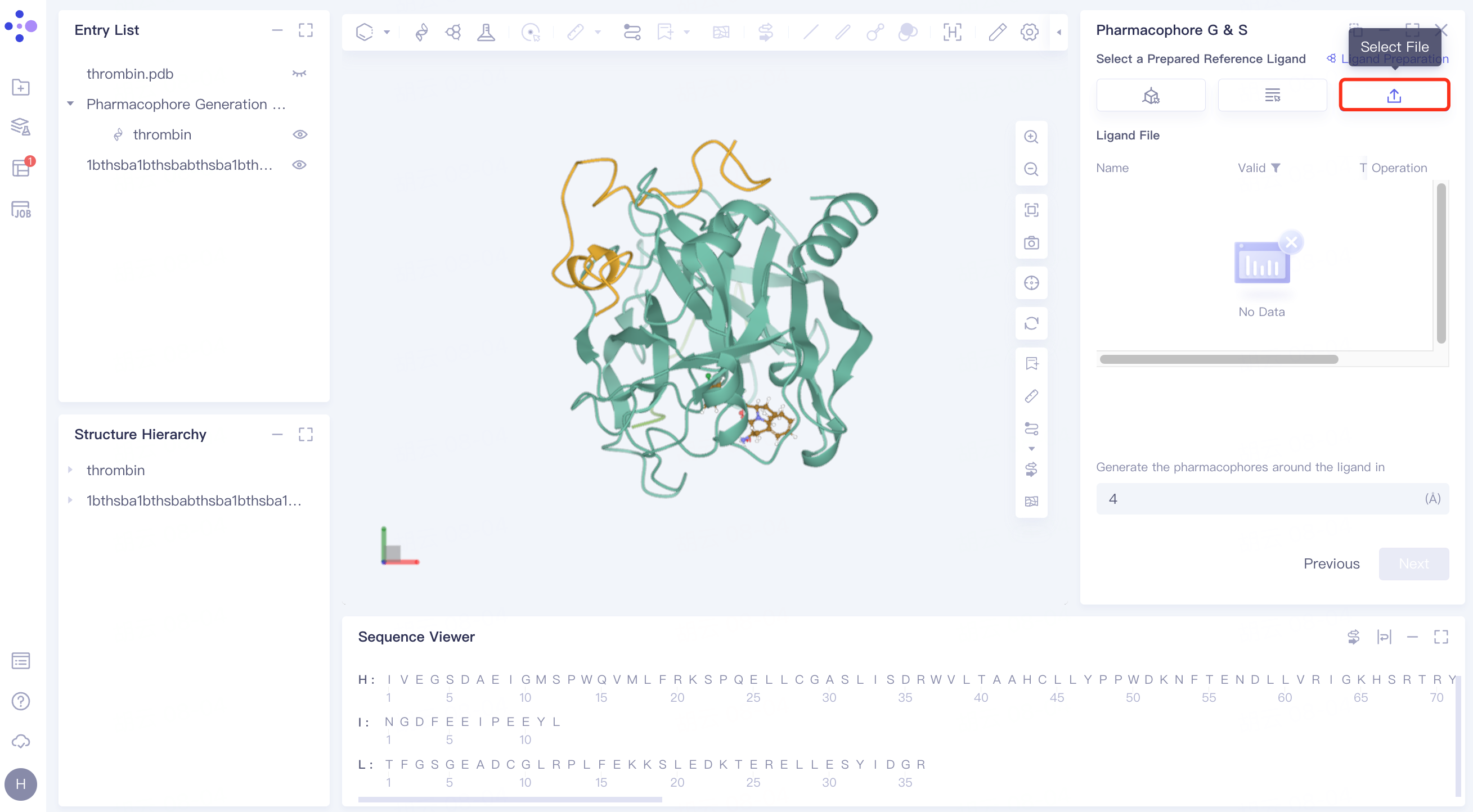
2.2.3 set pharmacophore generation parameters
Generate the pharmacophores around the ligand in ( ) Å: Sets the radius around the ligand to generate the pharmacophore, the default is 4.
Click "Next" to go to the next step.
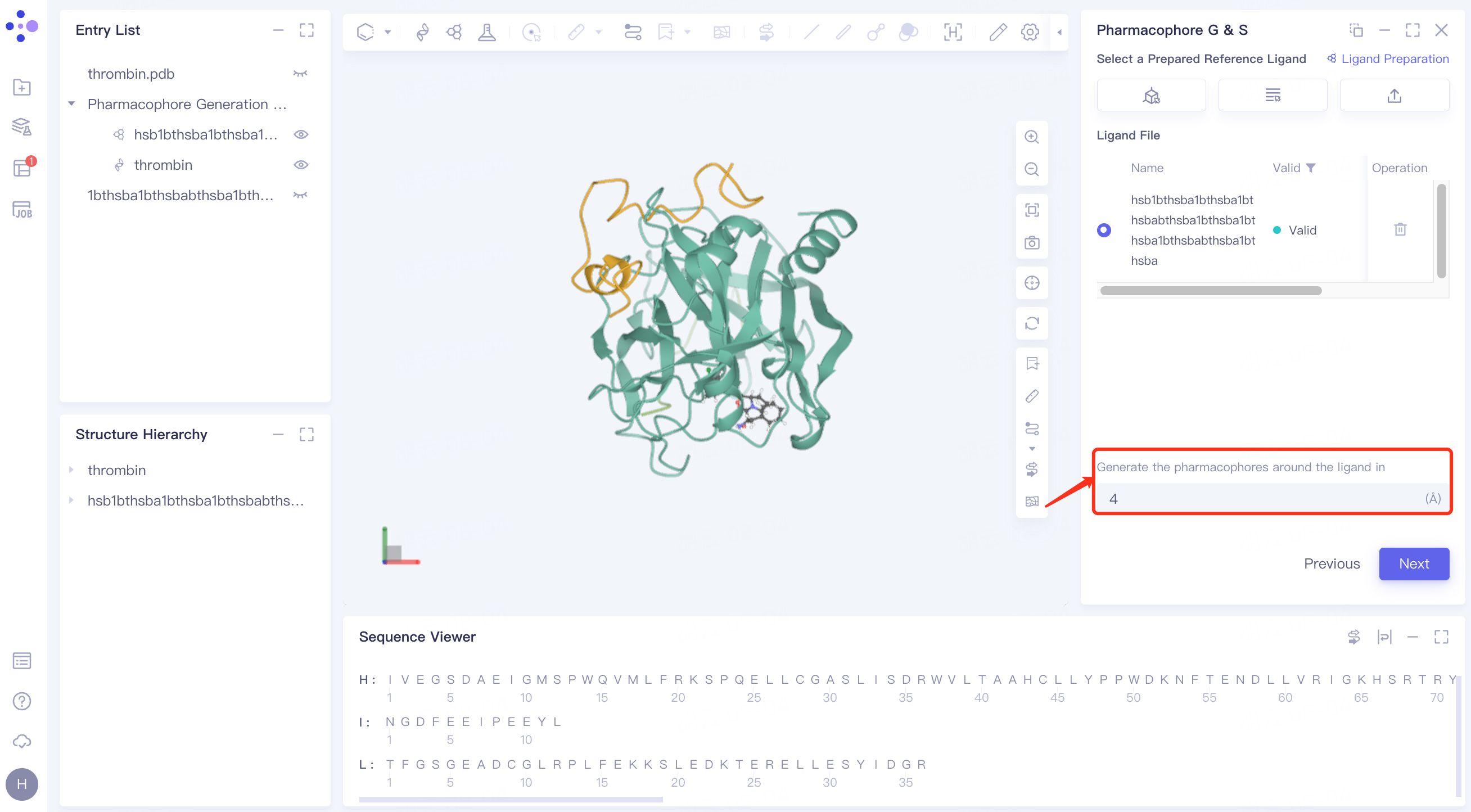
2.2.4 upload ligand
Steps are the same as 2.2.2.
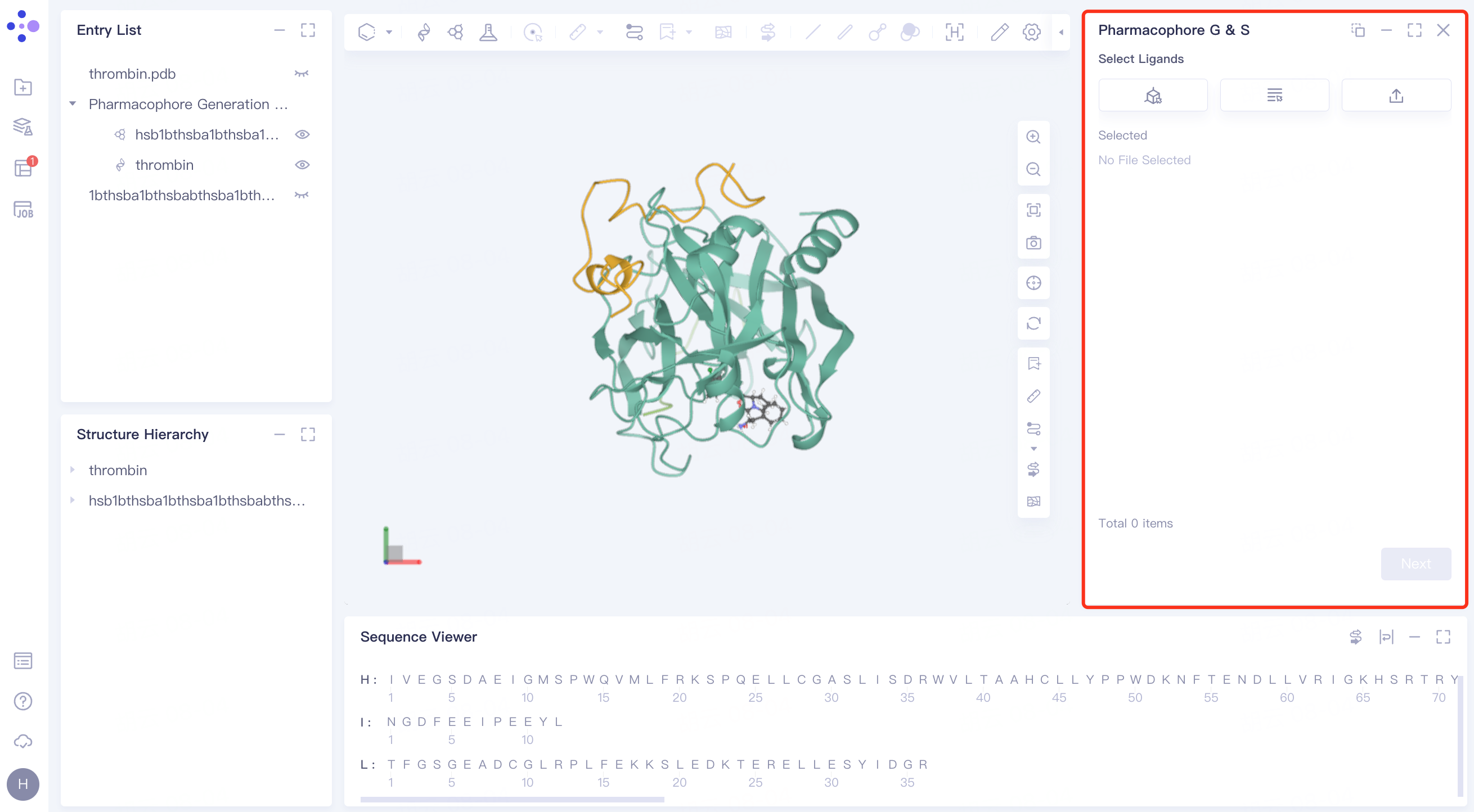
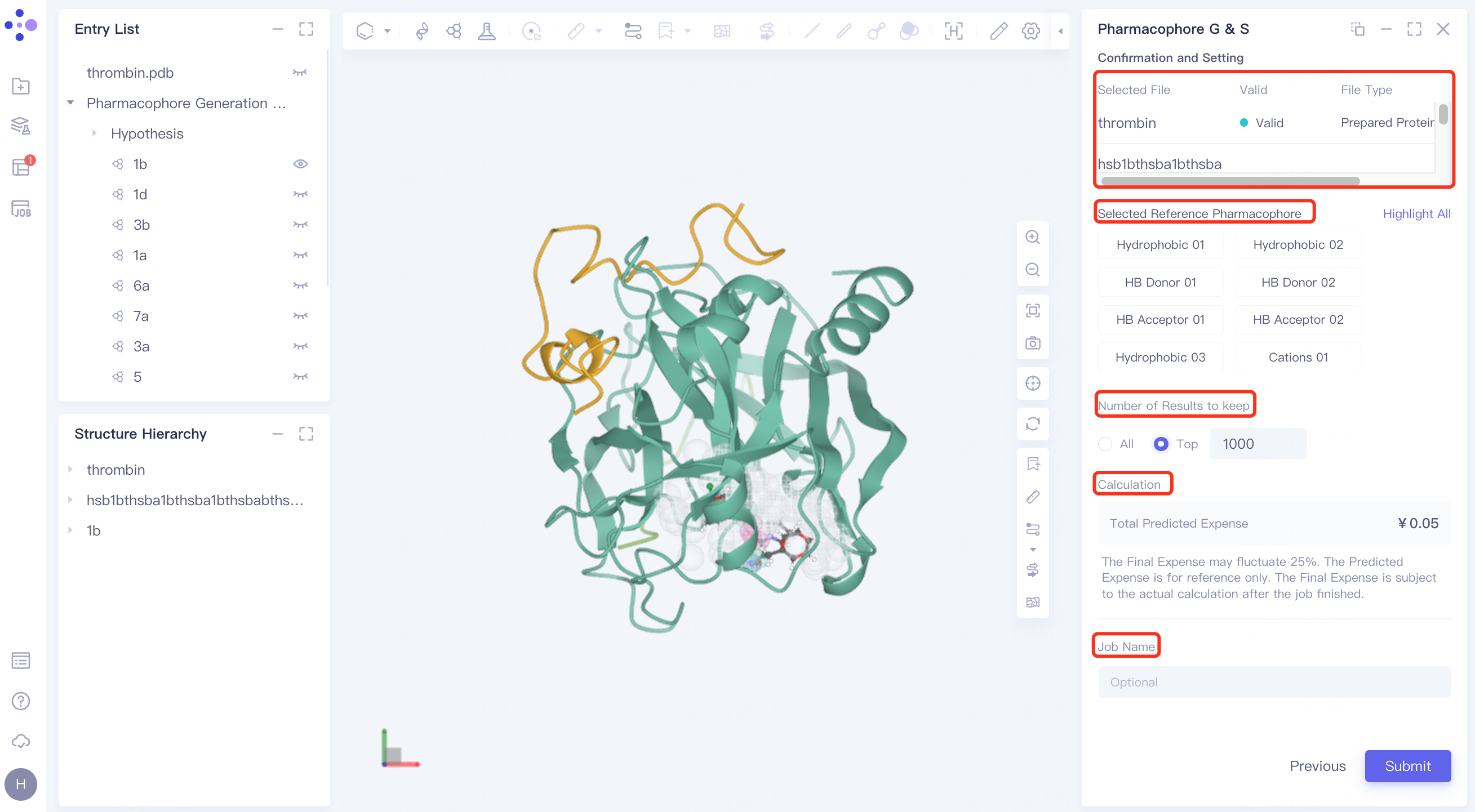
2.2.5 confirmation and parameter setting
Verify that the entered receptors, reference ligands, and ligand files required for screening are correct at the list.
Select Reference Pharmacophore: Click on the pharmacophore below to view the pharmacophore in the 3D Workspace window, the selection of pharmacophore is not supported here for the time being;
Number of Results to Keep: Optional All or Top (1~ 1000).
Calculation: Confirm the calculation cost of the task;
Name the task at "Job Name";
Click "Submit" to submit the task.
2.3 Receptor-free pharmacophore screening
2.3.1 selection of reference ligands and pharmacophore generation parameter settings
There are three ways to upload ligands, and the steps are the same as 2.2.2.
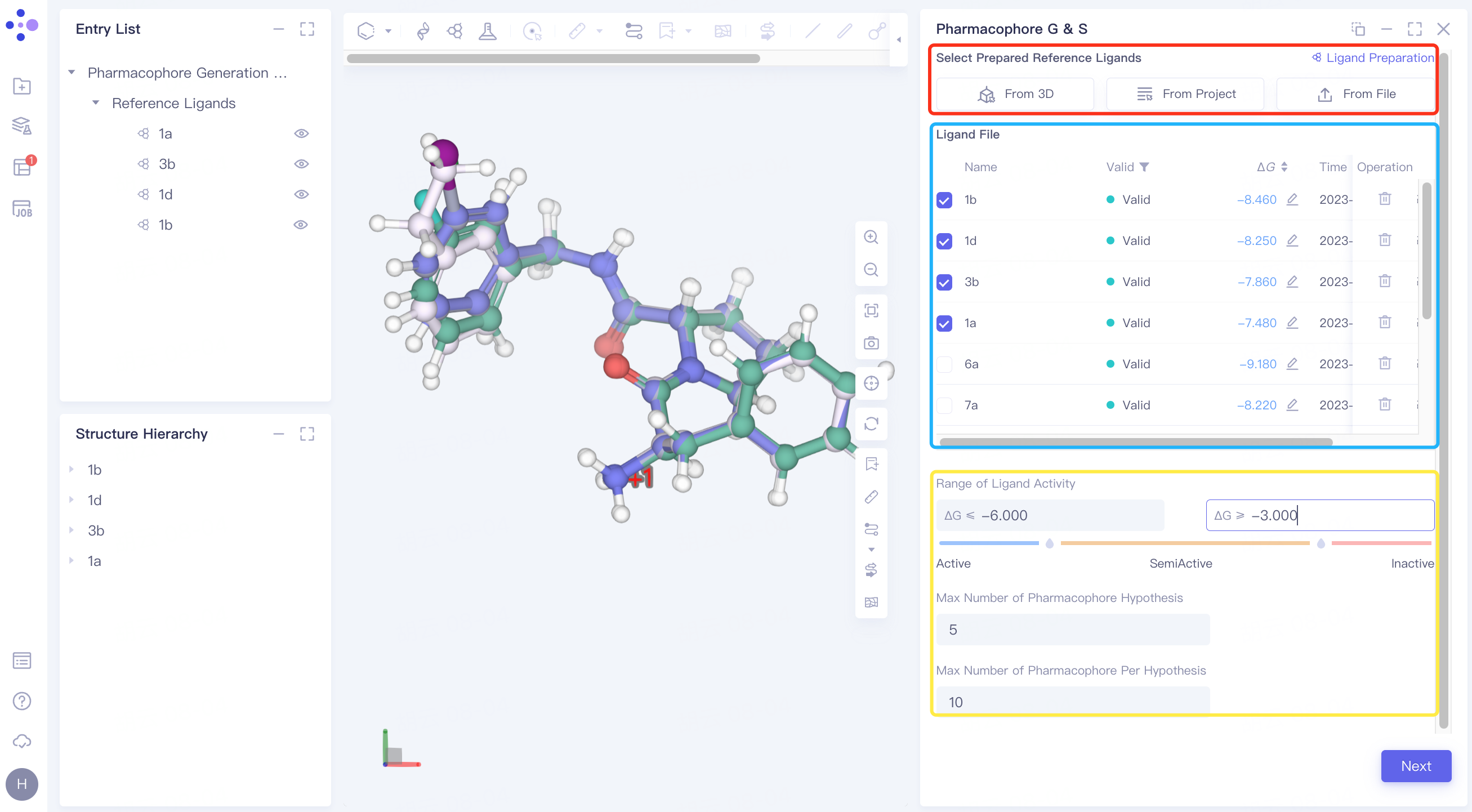
When the upload ligand is processed into the Valid state, n ligands (number of active ligands > = 2) are checked in the checkbox as reference ligands.
Range of Ligand Activity: The active/inactive range of the molecule is defined by ΔG;
Max Number of Pharmacophore Hypothesis: Set the maximum number of Pharmacophore model generation;
Max Number of Pharmacophore Per Hypothesis: Sets the maximum number of pharmacophore characteristics per pharmacophore model.
2.3.2 selection of ligands
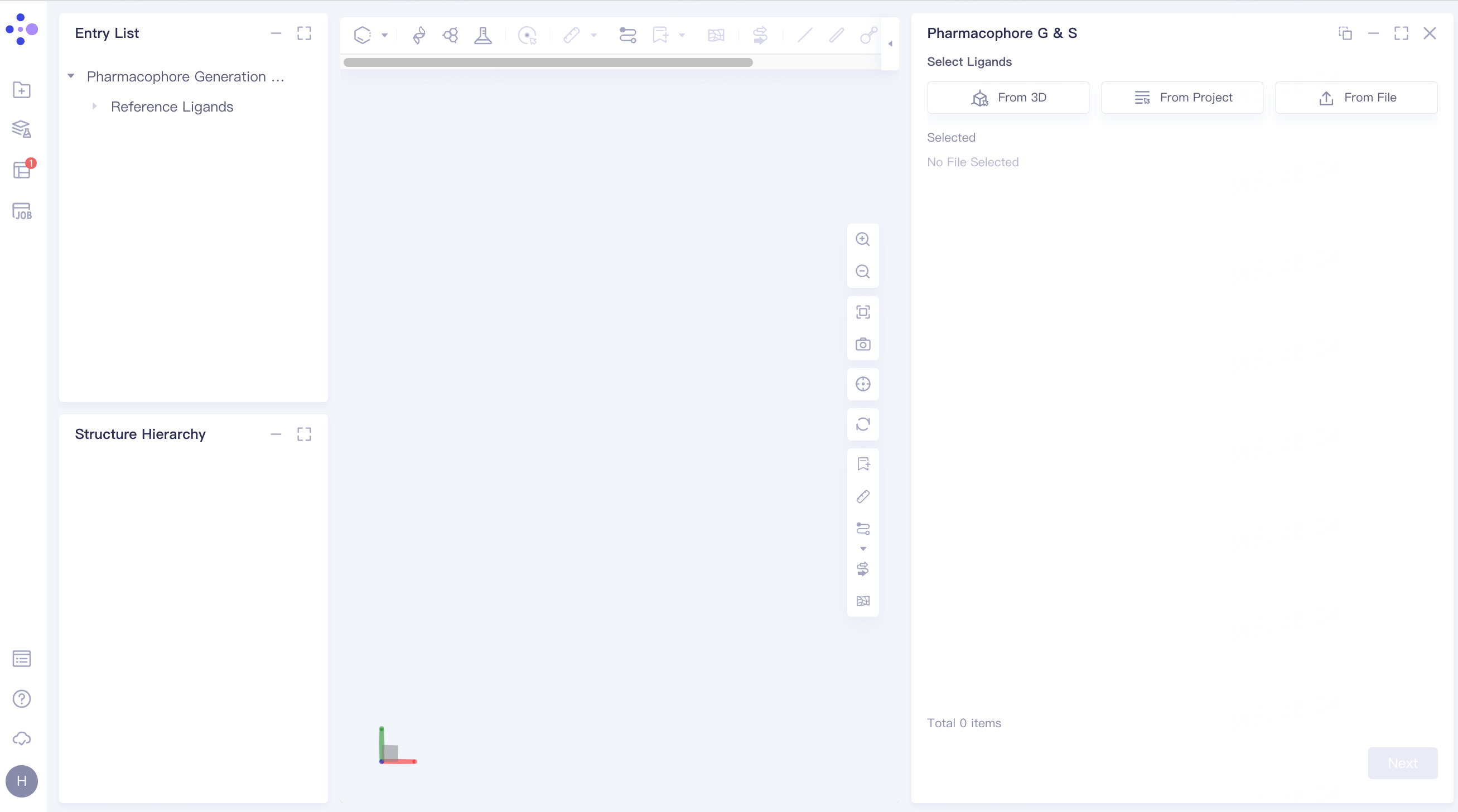
There are three ways to upload ligands, and the operation steps are the same as 2.2.2.
2.3.3 pharmacophore
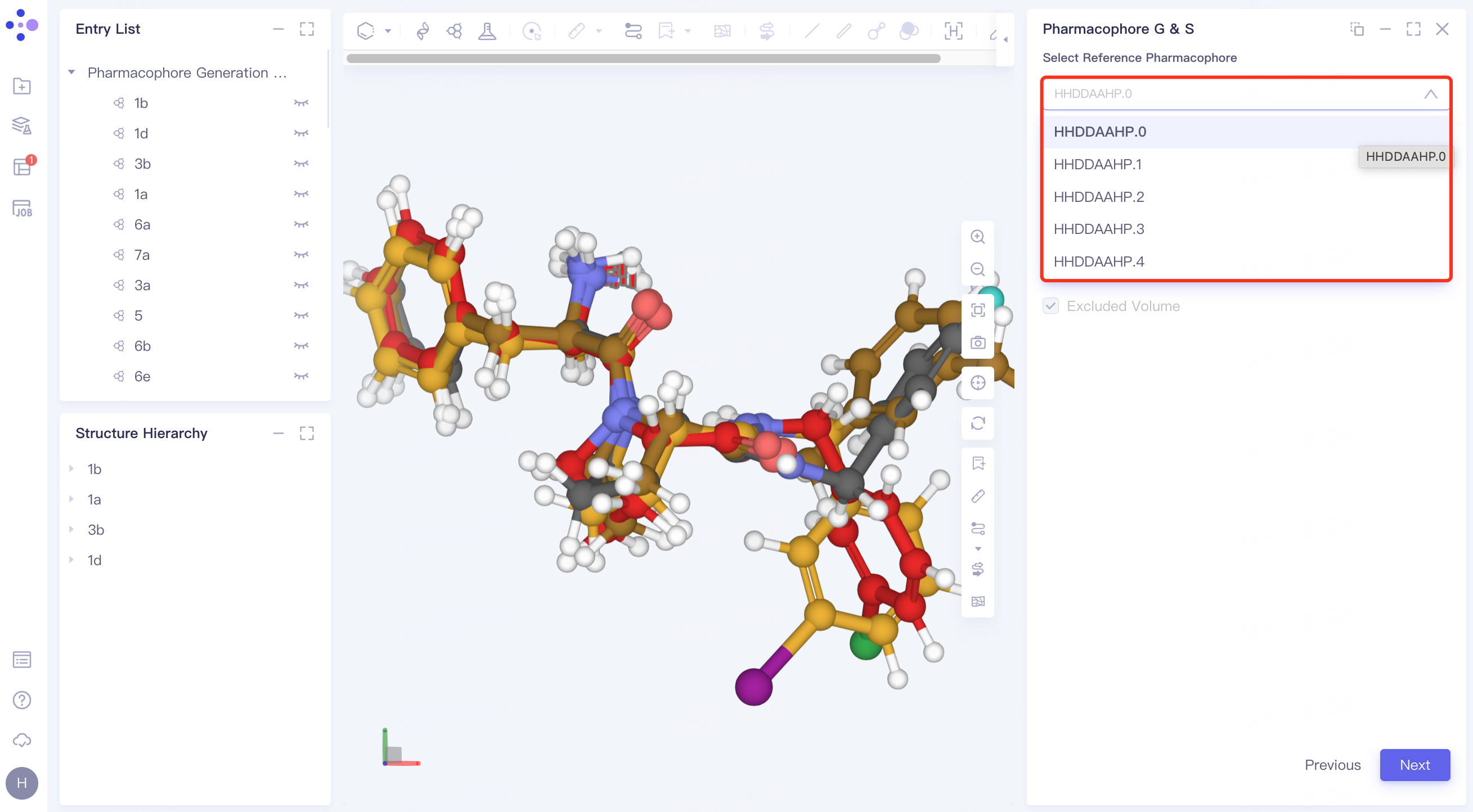
Select Reference Pharmacophore: Select Reference Pharmacophore from the drop-down window.
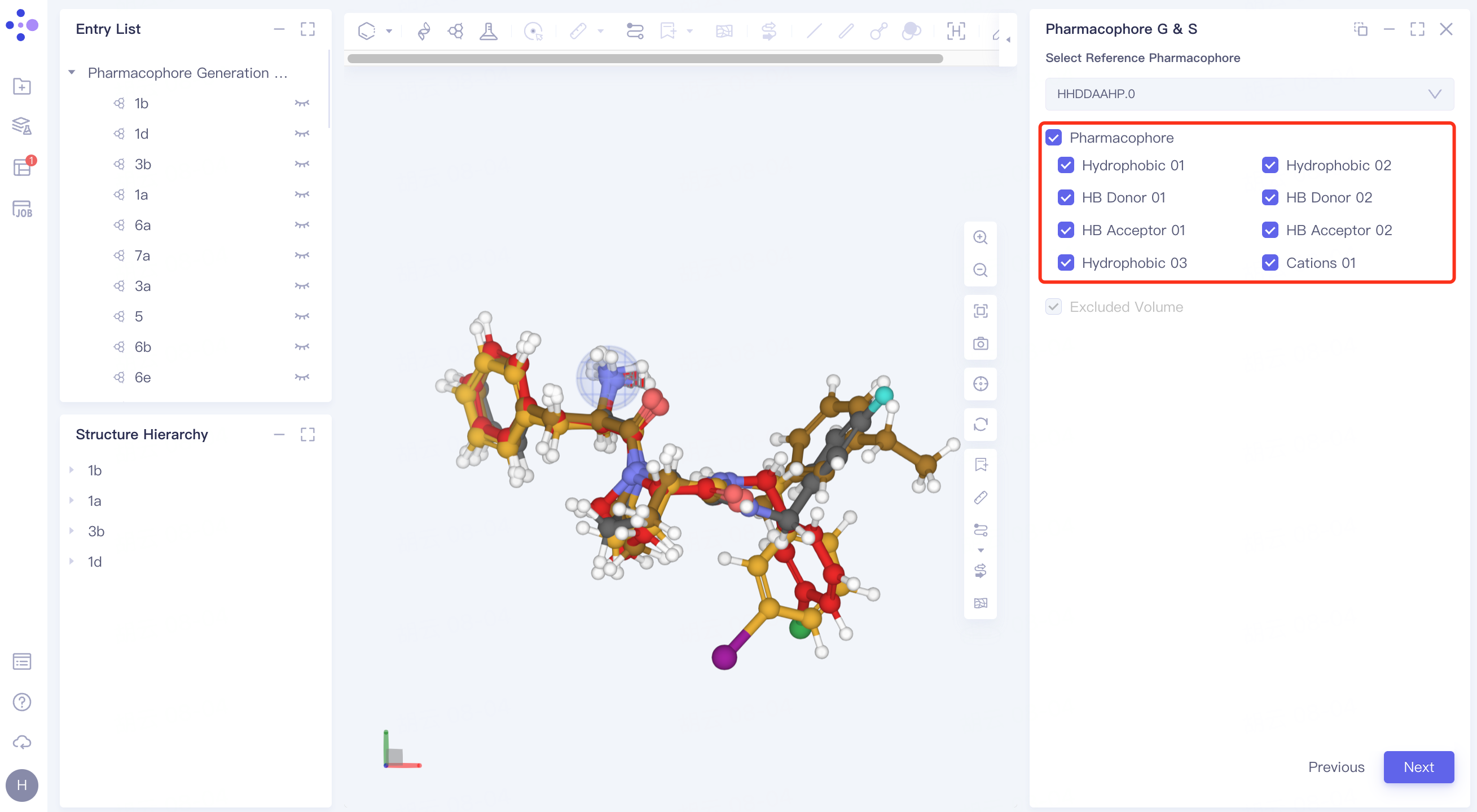
Adjust the pharmacophore under Pharmacophore.
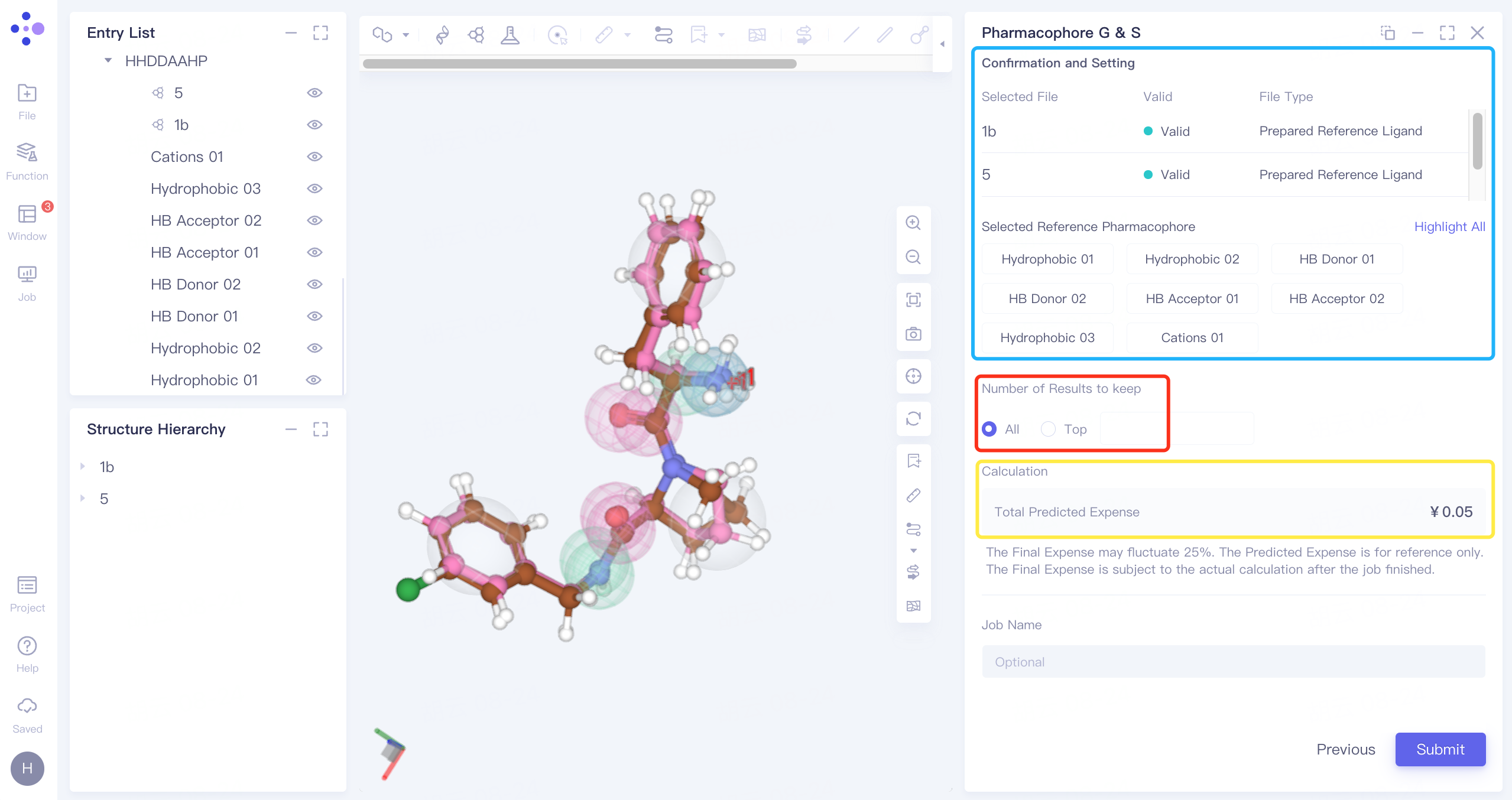
2.3.4 Confirmation and Setup
Basket: Confirm whether the uploaded reference ligands , ligands to be screened and pharmacophore are wrong;
Red box: Set the number of results to keep, and select "All" to keep all results;
Yellow box: confirm the calculation cost;
Name the task "Thrombin_2" at Job Name;
Click "Submit" to submit the task.
3. Results show
3.1 Entrance
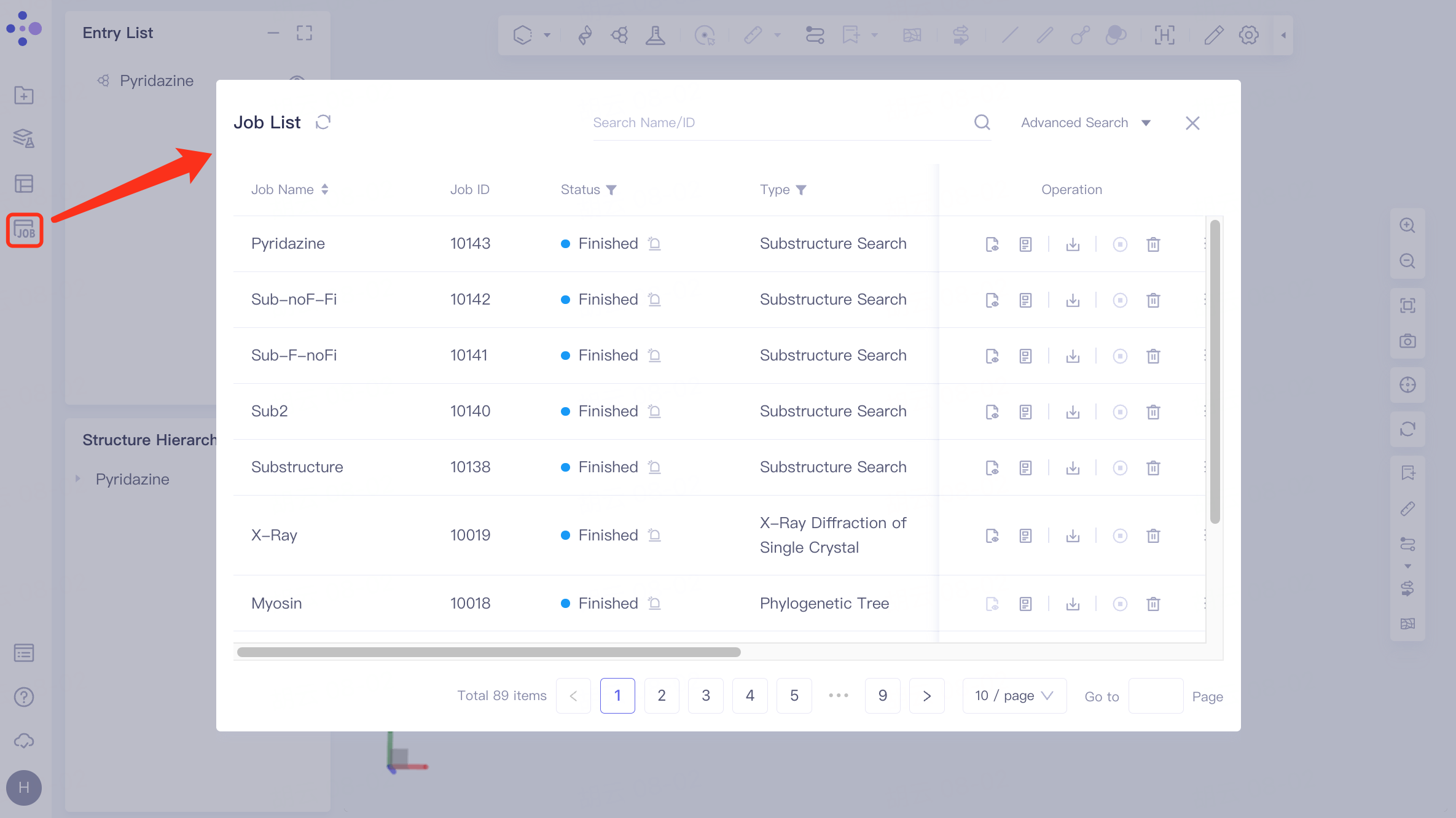
The left general menu bar Job → Find the required task.
The task can be found by searching for Job Name or by filtering through Job Type.
3.2 Pharmacophore results with receptor protein
3.2.1 Results
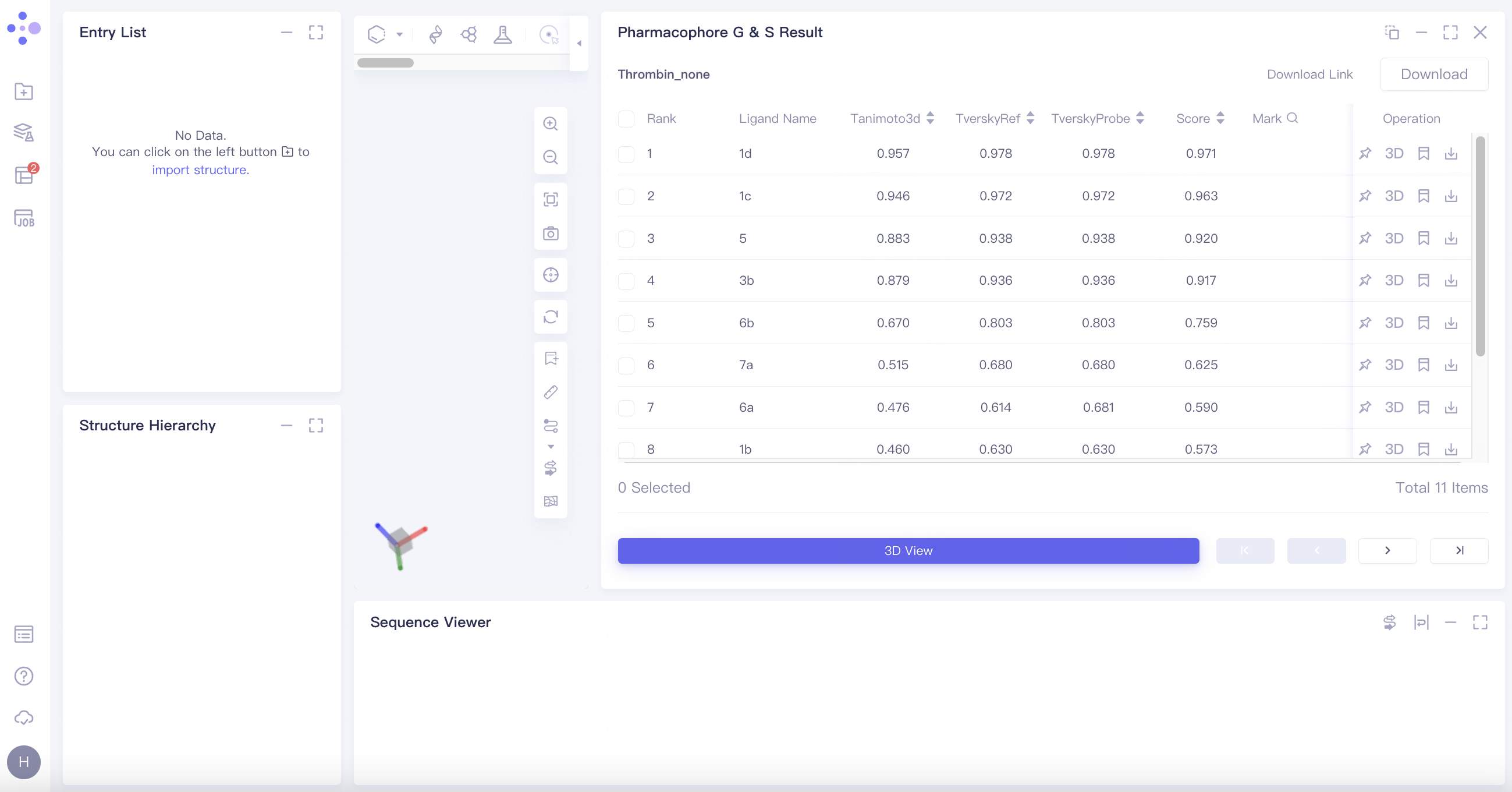
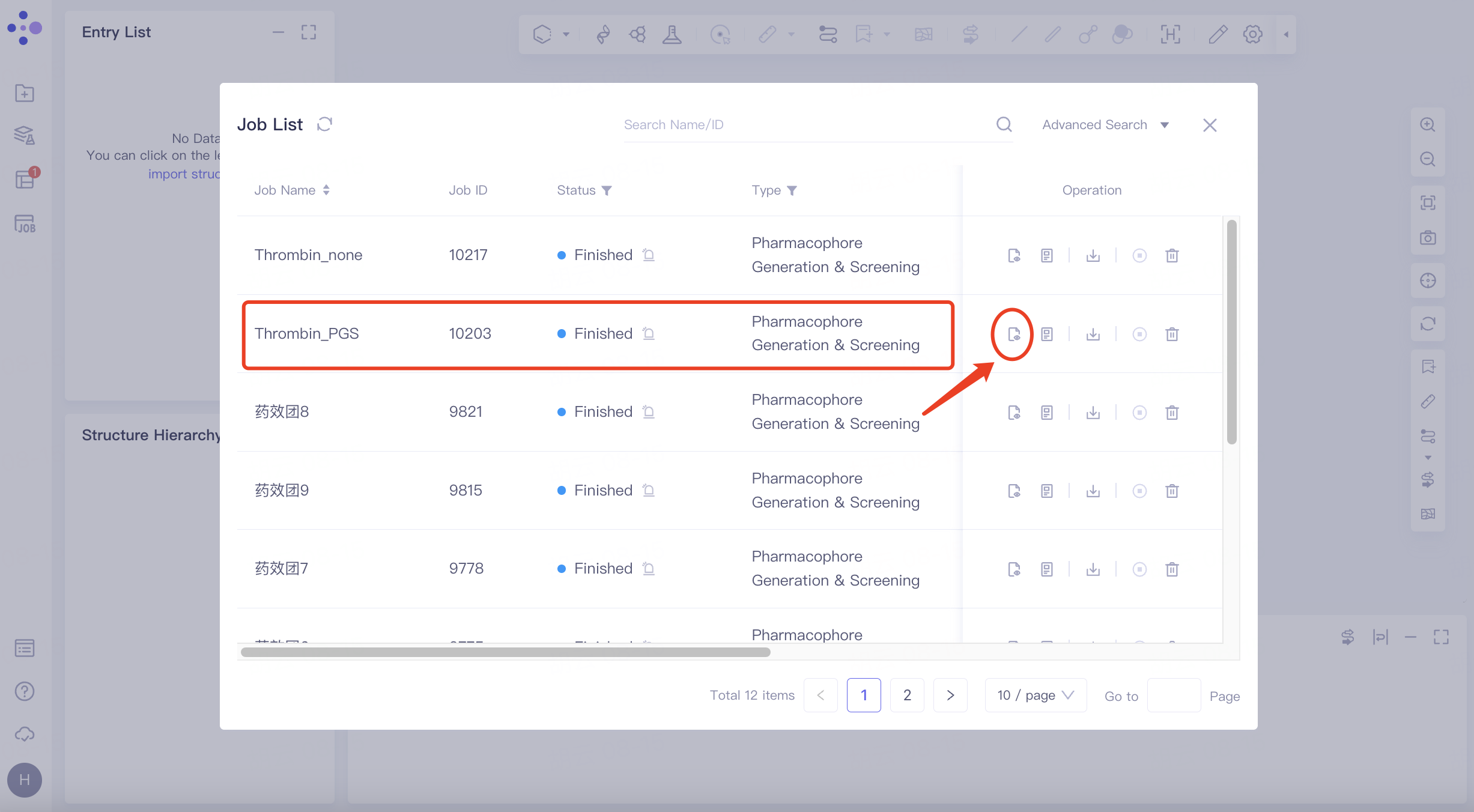
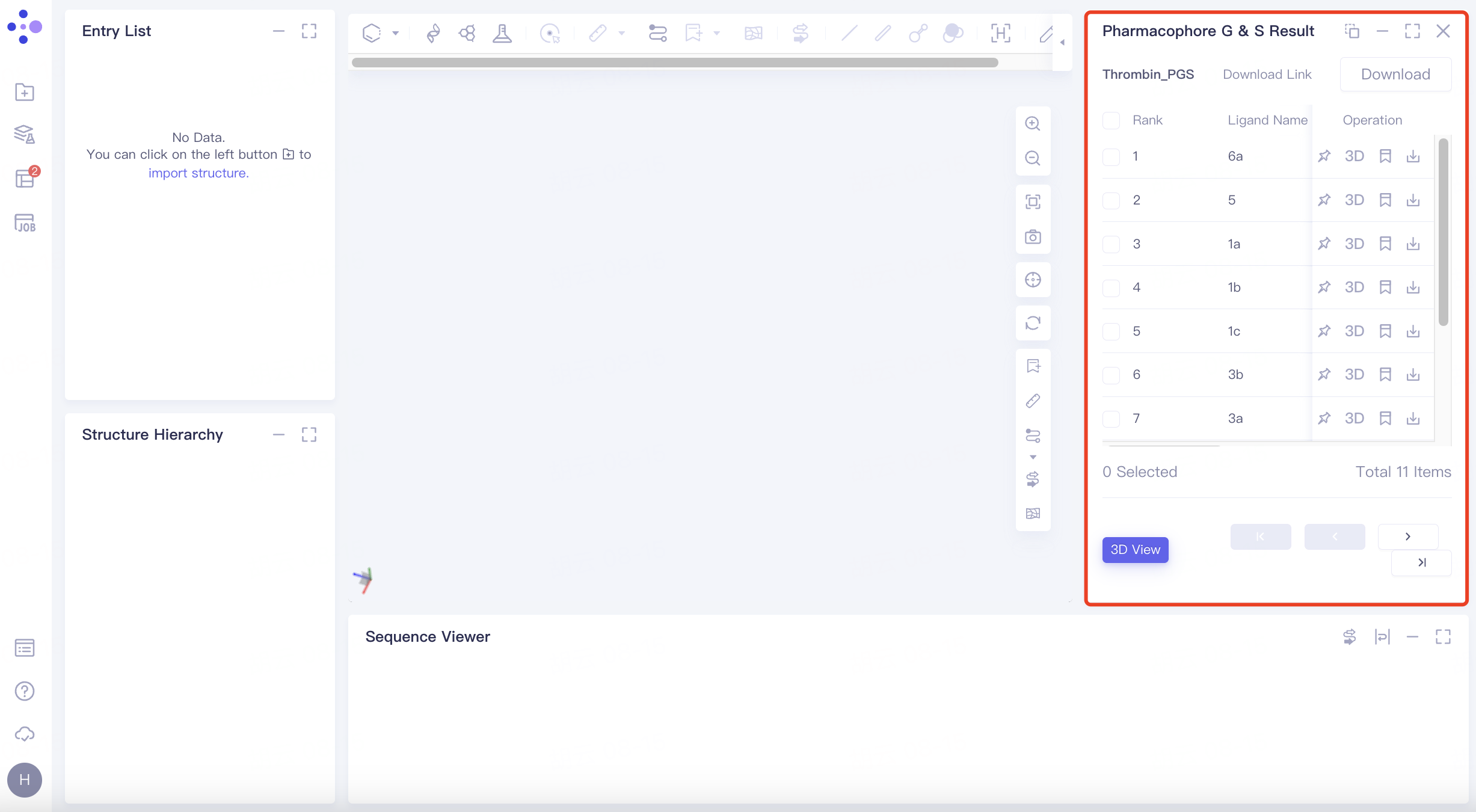
Select the task you want to view, and click Show in the Operation column to display the result of the task. The interface is shown in the figure.
 |  |
3.2.2 Results
Results table
The result list is sorted by the value of the scoring function Score by default;
The virtual screening of pharmacophore with receptor protein belongs to the virtual screening method based on the 3D shape of the ligand . By calculating the 3D similarity between two molecules, it is judged whether the molecule is active. Generally, the Tanimoto coefficient and the Tversky coefficient are used to measure the two molecules. The degree of 3D similarity between them, of which Tversky is one of the best similarity methods for similarity search and skeletal transition.
Scoring function (virtual screening method based on ligand 3D shape):
Tanimoto3d: Between the values [0, 1], the larger the match with the pharmacophore;
TverskyRef: Between the values [0,1], the larger the match with the pharmacophore;
TverskyProbe: Between the values [0, 1], the larger the match with the pharmacophore;
Score: The average of the above three scoring values.
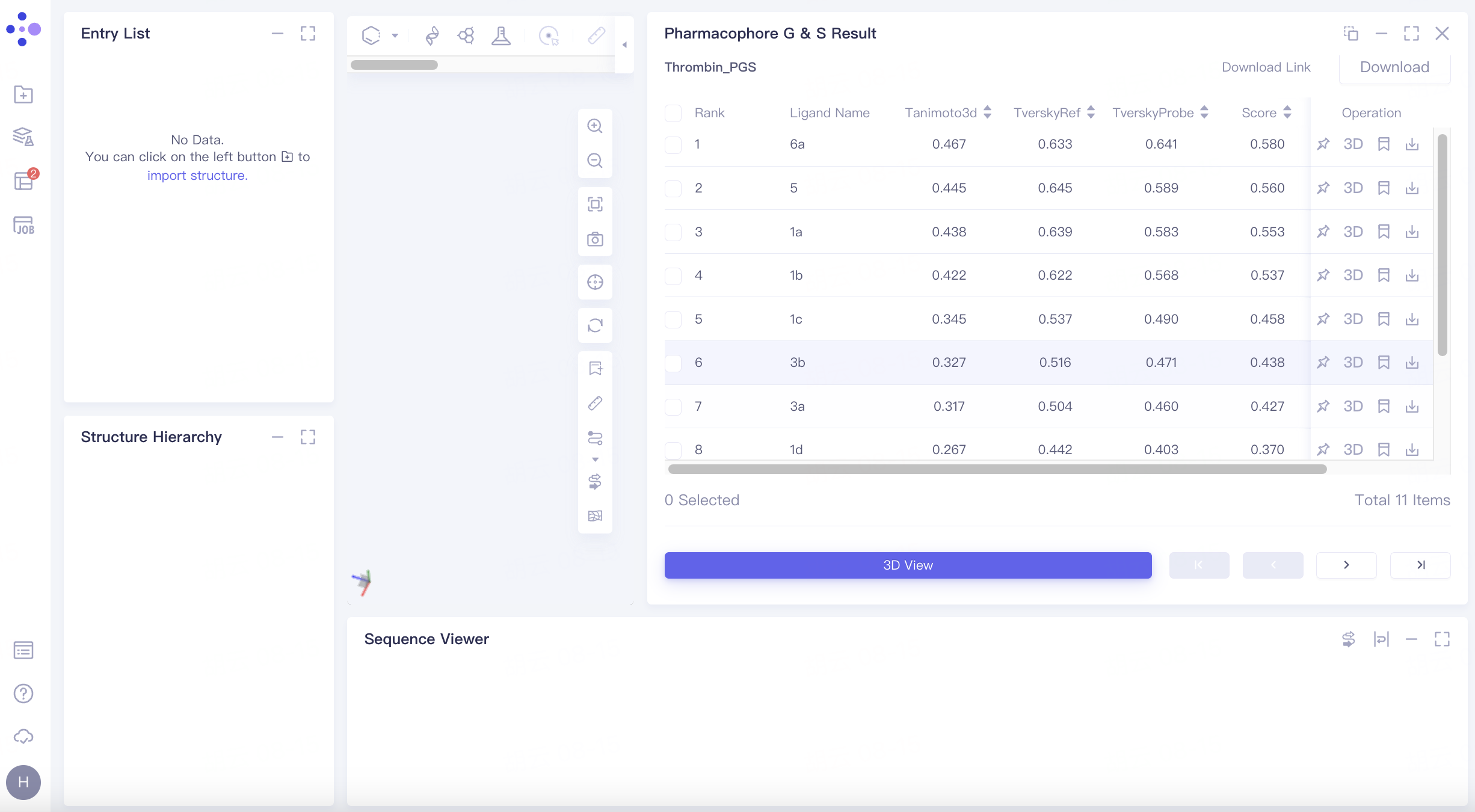
Operation
There are 4 operation options under Operation:
Fix: Fix the display in 3D Workspace.
Show in 3D Workspace: Displays the ligand in the 3D Workspace interface.
Mark: Click "Mark" to manually label a ligand. After labeling, it will be displayed in the Mark column, and search by label is available.
Download: Download the docked molecule, which can be saved in .sdf and .mol formats.
3D View: Ligand can be displayed in batches as follows:
Select the ligand molecule to be displayed in the checkbox → click "3D View" → the receptor and ligand are displayed in the 3D Workspace window.
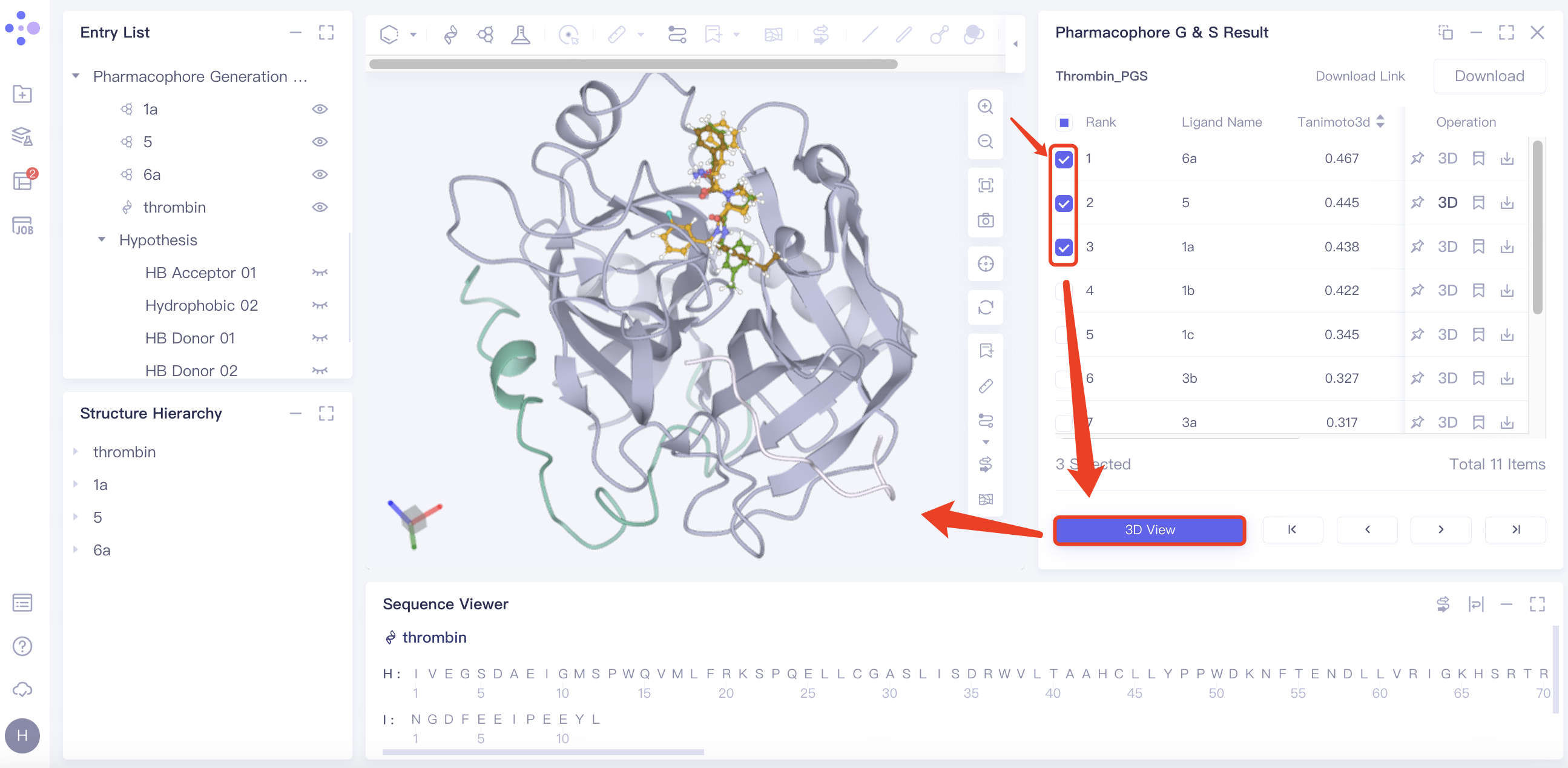
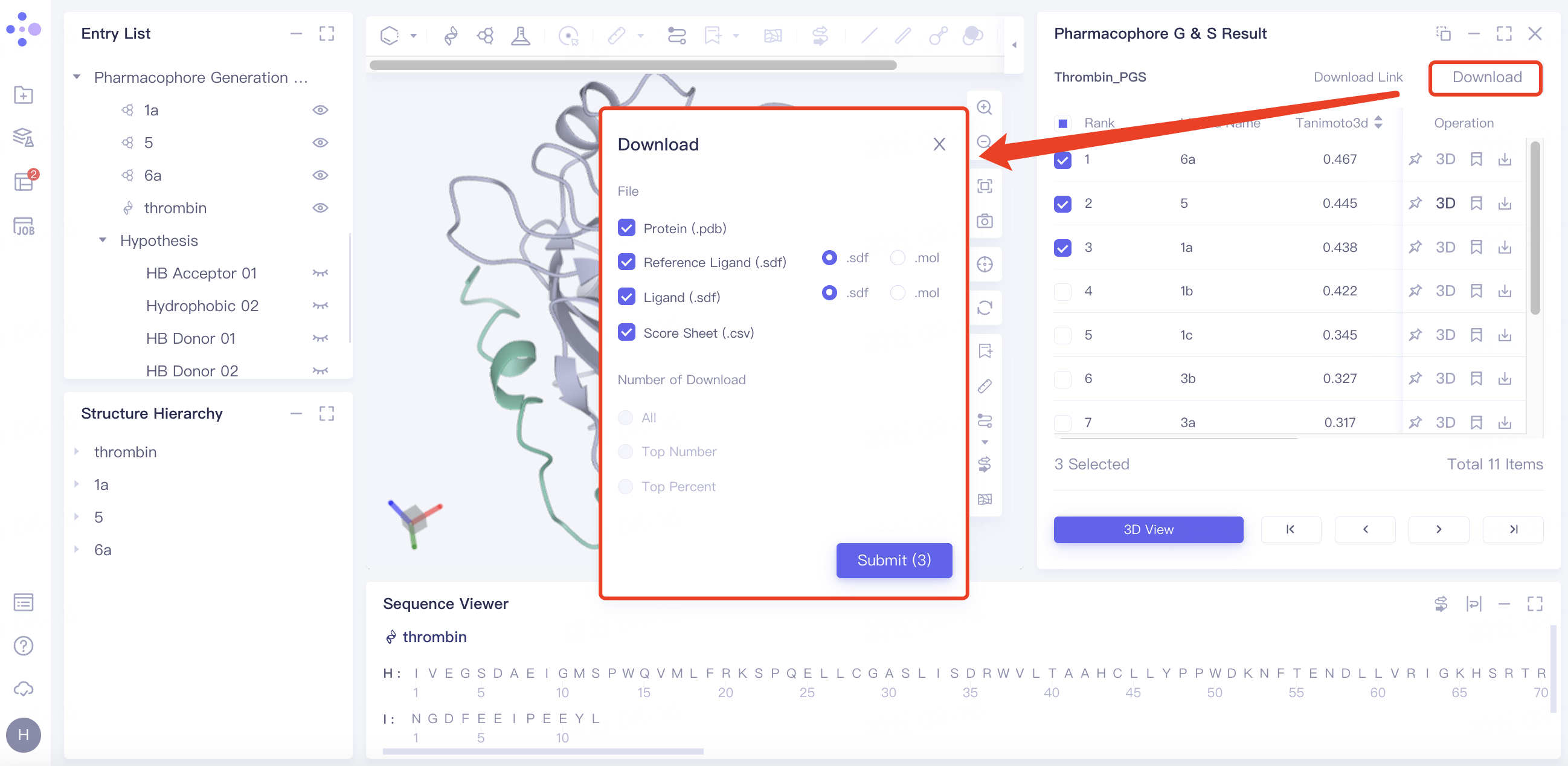
Batch download pharmacophore results:
Select the desired ligand in the checkbox → click "Download" in the upper right corner → select the content to be downloaded in the pop-up download window → click "Submit".
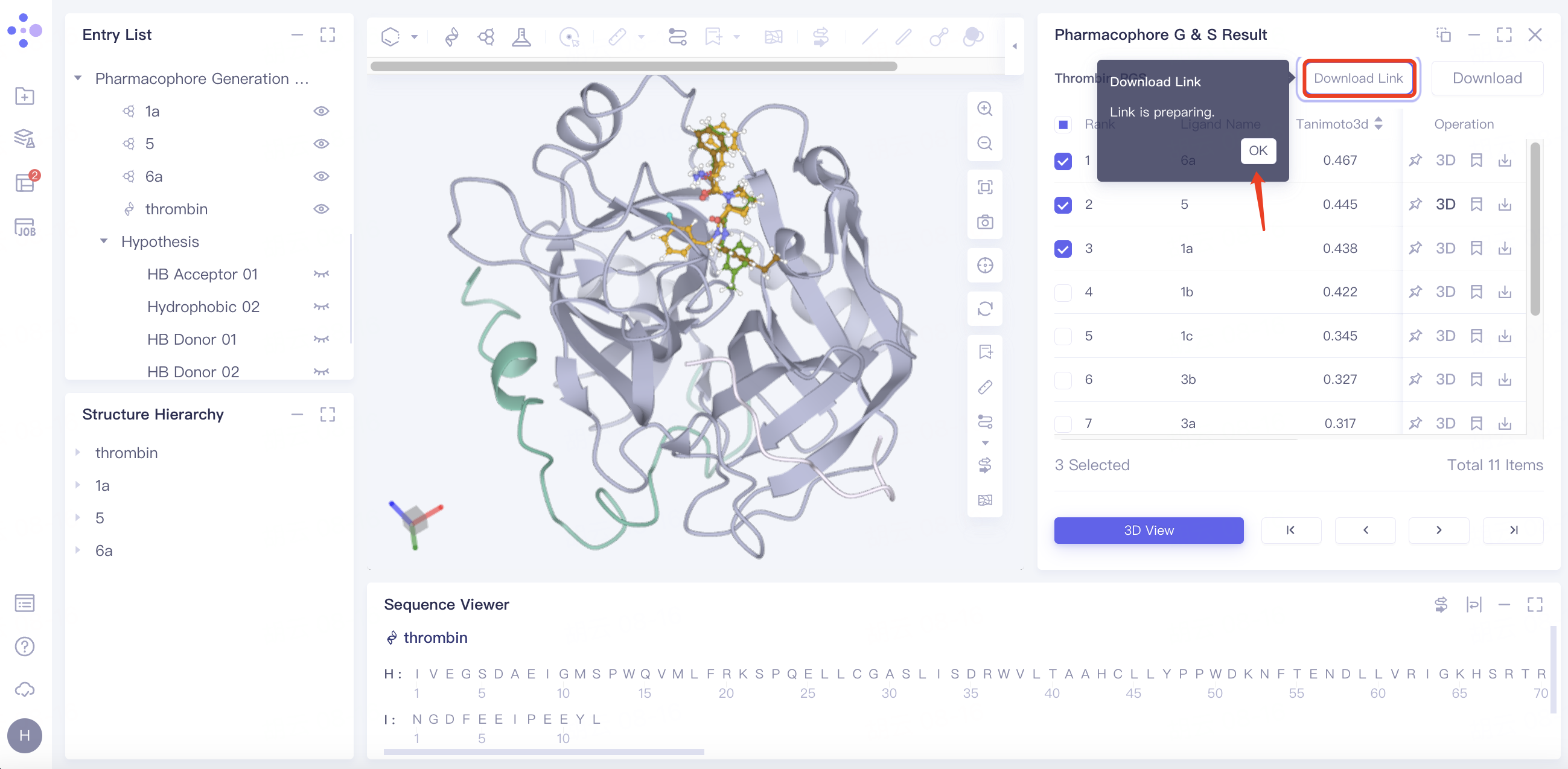
"Download Link" flashes and displays "Link is preparing", click "OK".
Click "Download Link" and click the link under URL in the pop-up window to start the download.
 | 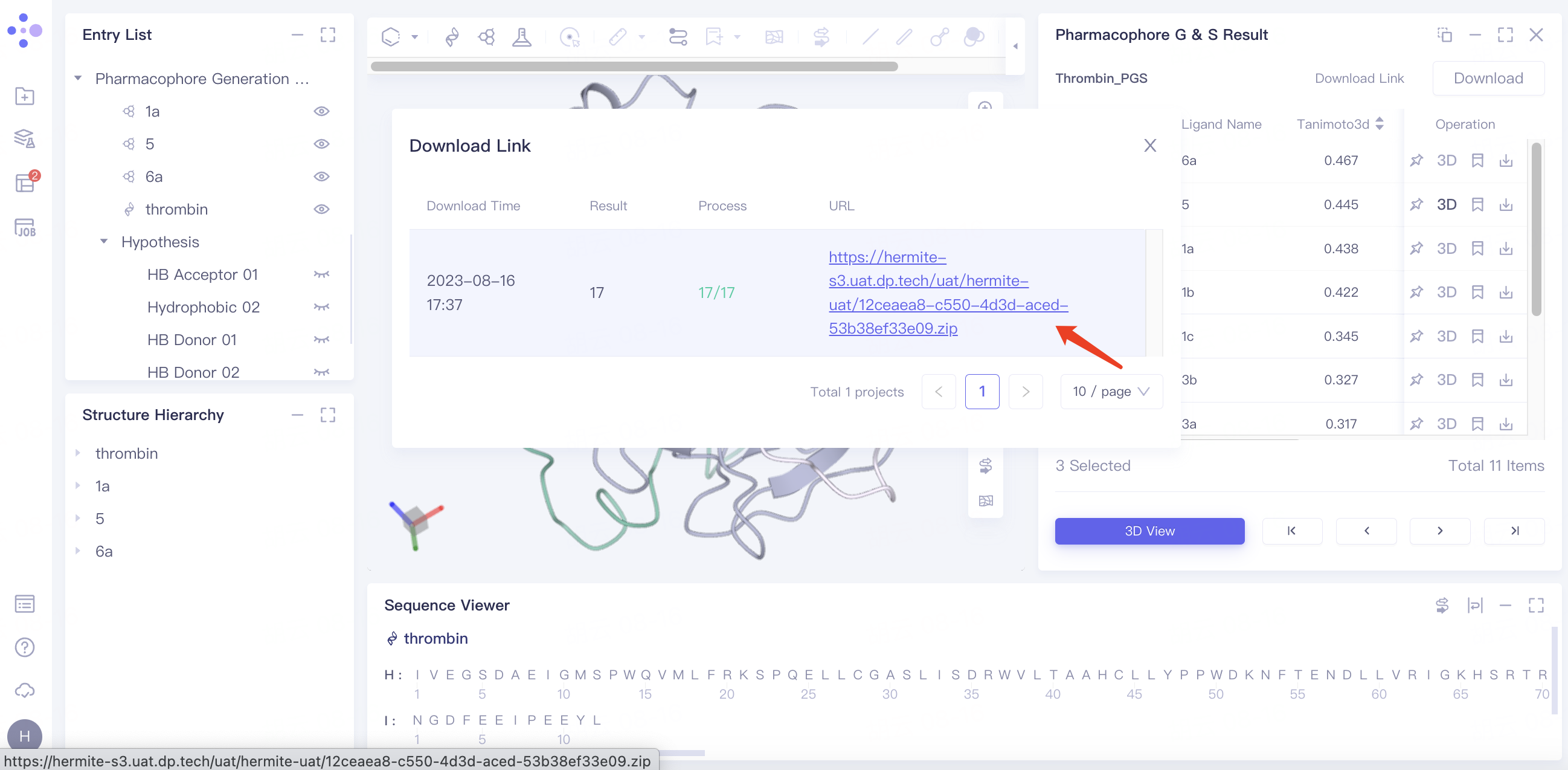 |
3.3 Pharmacophore results for receptor-free proteins
3.2.1 Results
Select the task you want to view, and click Show in the Operation column to display the result of the task. The interface is shown in the figure.
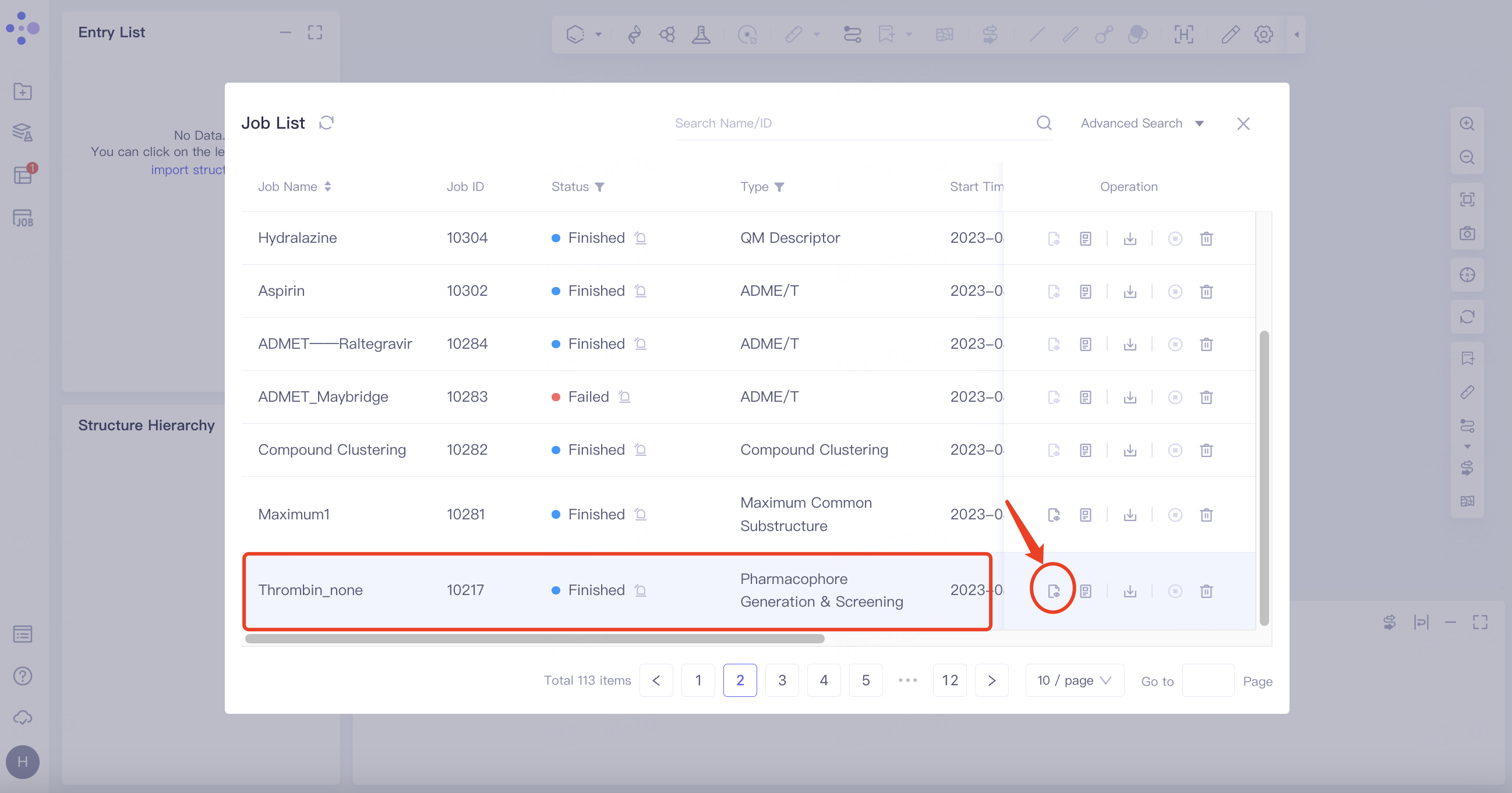 | 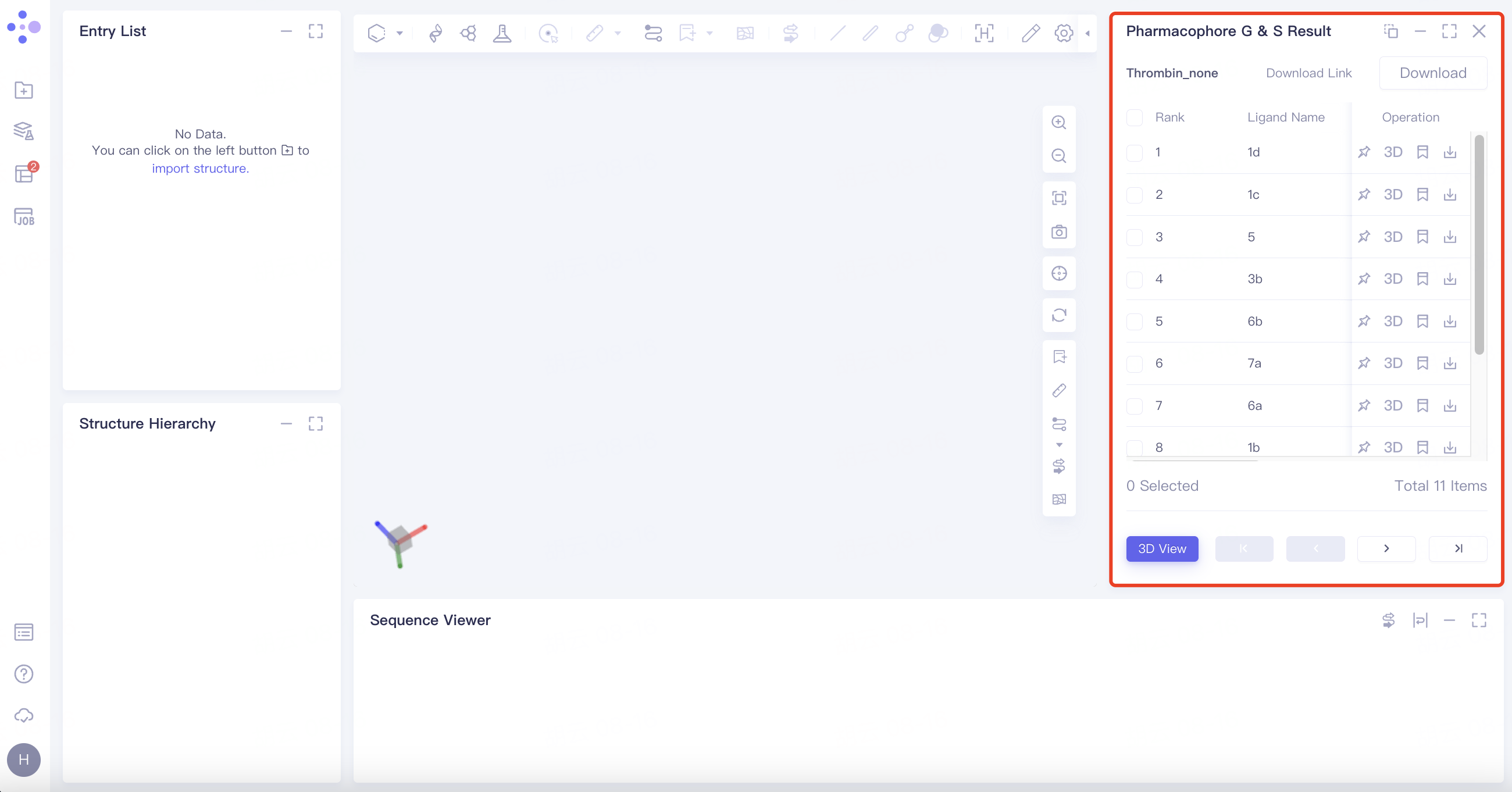 |
3.2.2 Results
The Same with 3.2.1.